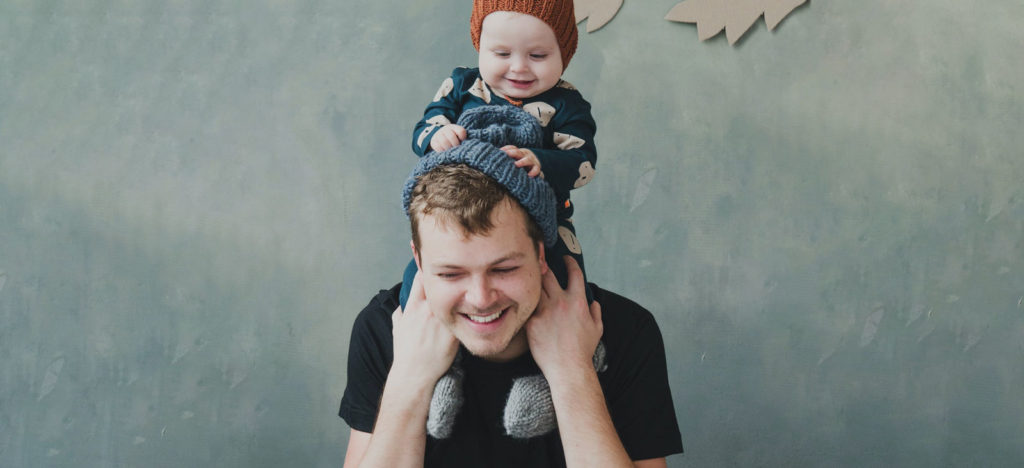
Our science
How our clinical team turns thousands of peer reviewed papers and scientific research into your personalised insights.

Eliminating guesswork through science
When Associate Professor Les Sheffield founded myDNA in 2007, he had one vision in mind. His quest was to use science to eliminate the guesswork and trial-and-error from prescribing medication. Backed by decades of scientific experience as a clinical geneticist, he has since been able to help thousands of people.
Since then, myDNA has expanded into helping people eliminate the guesswork from other aspects of their health and wellness. With the research always evolving, we’re commited to making sure all our recommendations are based in science. Our heritage is in the science, and it’s at the core of what we do every day.
Turning DNA insights into action
We want to empower the world to live healthier and happier lives.
By leveraging the world’s leading genotyping technology and scientific algorithms, this goal becomes a reality.
DNA Profile
Lifestyle inputs
Unique plans
Our scientific team does the hard work for you, decoding the secrets of your DNA and letting you know how you and your genetics can work together. We’ve collaborated with nutrition and fitness experts so you can improve your health with confidence knowing you’re backed by science.
We give you easy-to-understand, relevant and actionable plans that help you turn your DNA insights into action.
Our team of clinical experts are backed by the world’s leading research
120+ years of combined experience
100+ papers published
5000+ research publications reviewed
5 myDNA clinical studies conducted

Associate Professor Les Sheffield
Founder, Clinical Geneticist & Medical Director
Les has been at the forefront of genetic research since the 1980s. As a Clinical Geneticist based at Royal Women’s and Children’s Hospitals, he authored more than 100 scientific publications. A leader in his field, he founded myDNA in 2007 with the quest to eradicate guesswork of prescribing medication. To this day he continues to drive the development of new services as Medical Director.

99.9% accurate open-array technology
All recommendations backed by high quality science
We only use the highest quality research after the most rigorous review of scientific literature.
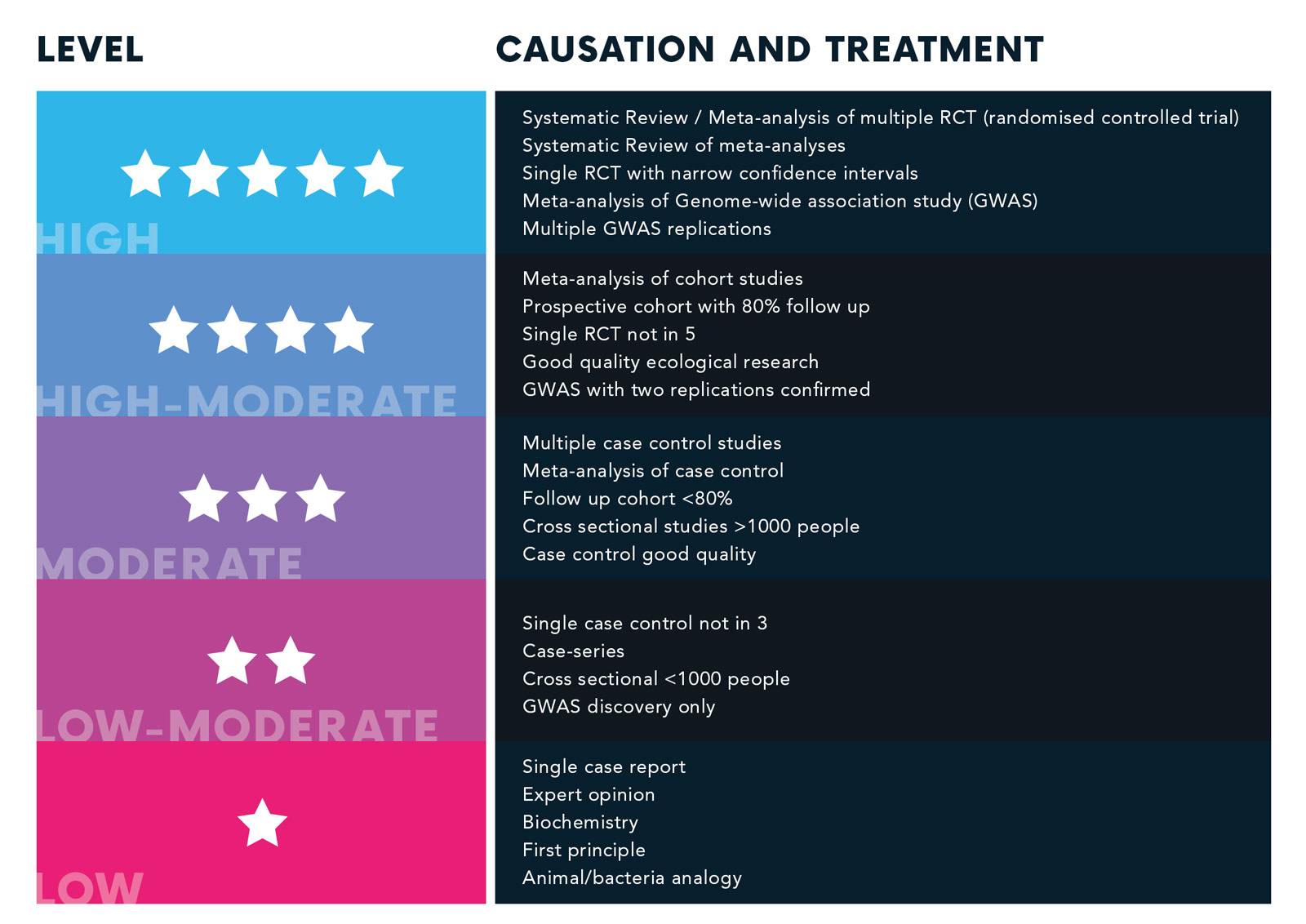
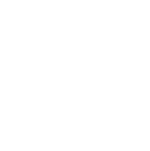
To make sure our recommendations are air-tight, we’ve developed our own evidence rating system. We’ve followed the guidelines set by the Oxford Centre of Evidence Based Medicine, meaning we’re up there with the best when it comes to scientific evidence.
Each piece of research we use to make your plans needs to achieve at least a 3 star rating.