Pharmacogenomics Testing for prescription medications
Multiple Category Medication Test
Multiple Category Kit

Select the Multiple Categories Test to discover your response to a range of medications, including mental health, pain, gastrointestinal, cardiovascular medications + more.
$198.00
Mental Health Medication Test
Single Category Kit

The Mental Health Test Kit will help inform the treatment of depression, anxiety, ADHD and other psychiatric conditions.
$149.00
Visit a supporting pharmacy or call us on 1300 436 373 to learn more and find out if your medication is covered.
Please note: results will not be released until the consultation with your healthcare practitioner has been undertaken.
Multiple Category Medication Test Kit
Understand how your DNA may affect your response to certain medications.

Discover more about how your DNA may affect your response to a range of medications, including mental health, pain, gastrointestinal, cardiovascular medications + more.
Results have a lifetime value. By purchasing a myDNA test, you and your doctor will be more informed to tailor your medications to your genetic profile.
Please note: results will not be released until the consultation with your healthcare practitioner has been undertaken.
$198.00


Mental Health Medication Test Kit
Understand how your DNA may affect your response to certain mental health medications.

The Mental Health Test Kit will help inform the treatment of depression, anxiety, ADHD and other psychiatric conditions.
Results have a lifetime value. By purchasing a myDNA test, you and your doctor will be more informed to tailor your medications to your genetic profile.
Please note: results will not be released until the consultation with your healthcare practitioner has been undertaken.
$149.00
Mental Health Medications
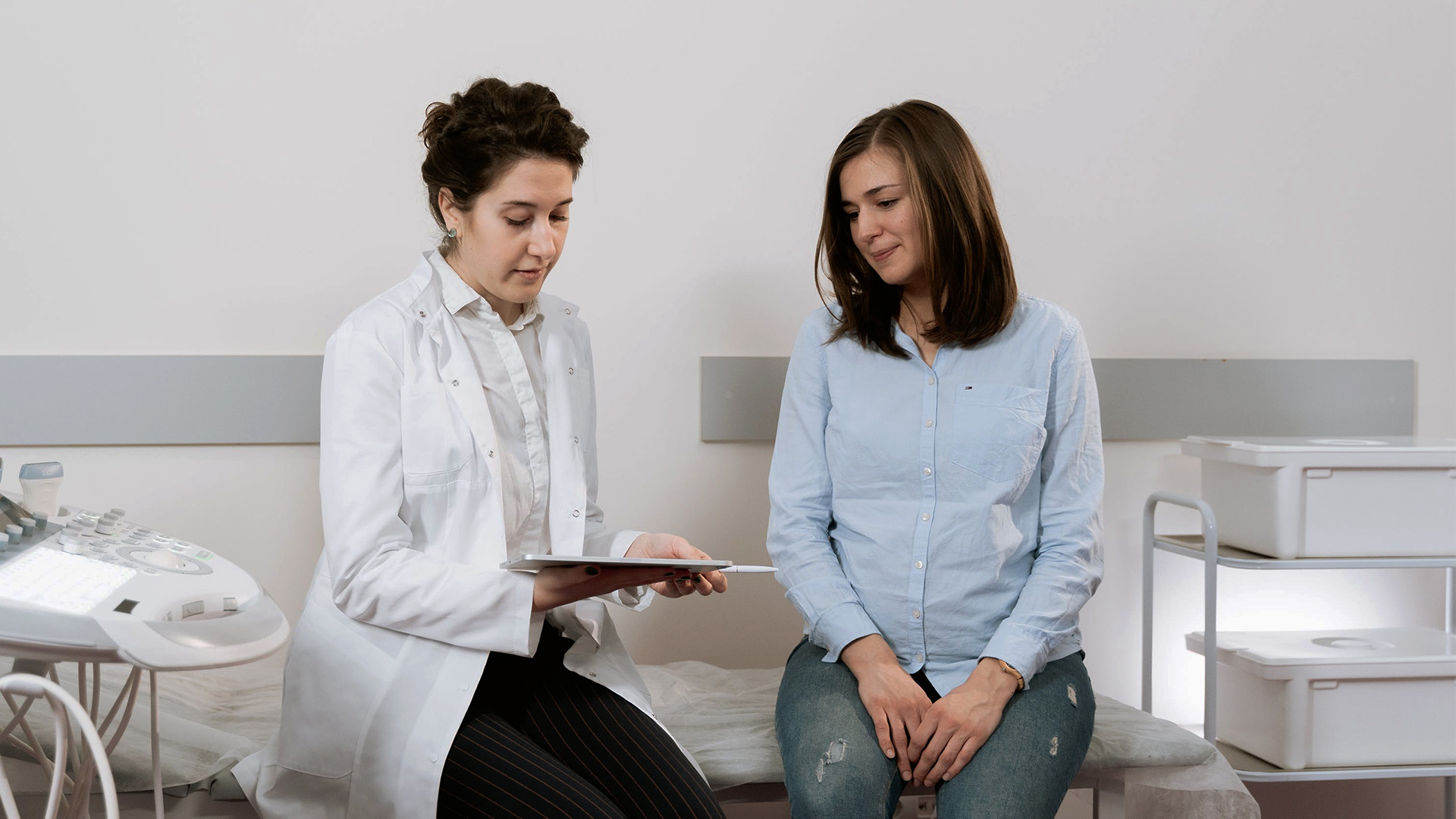

Pain Medications
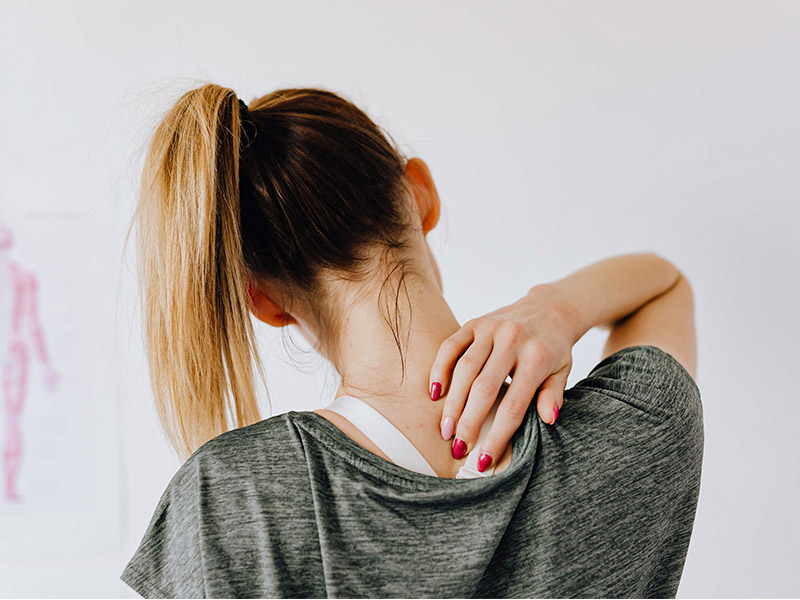

Gastrointestinal Medications
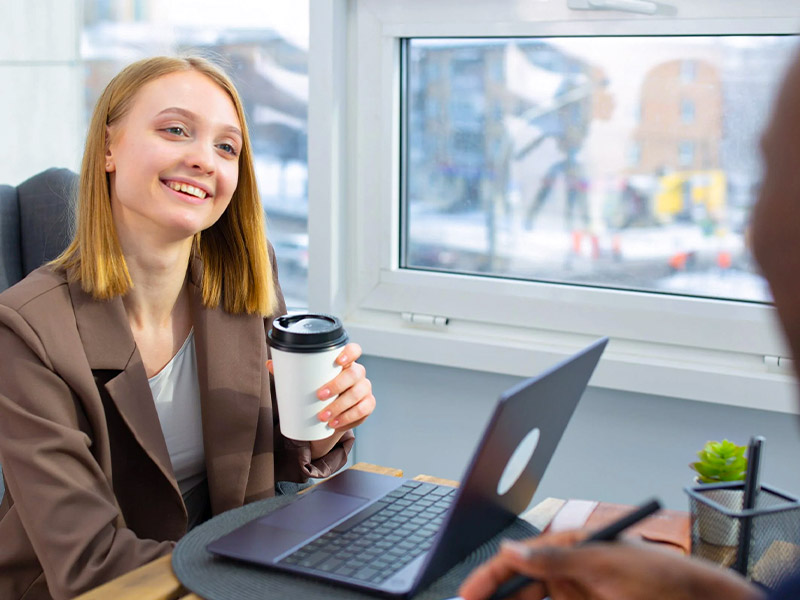

Cardiovascular Medications
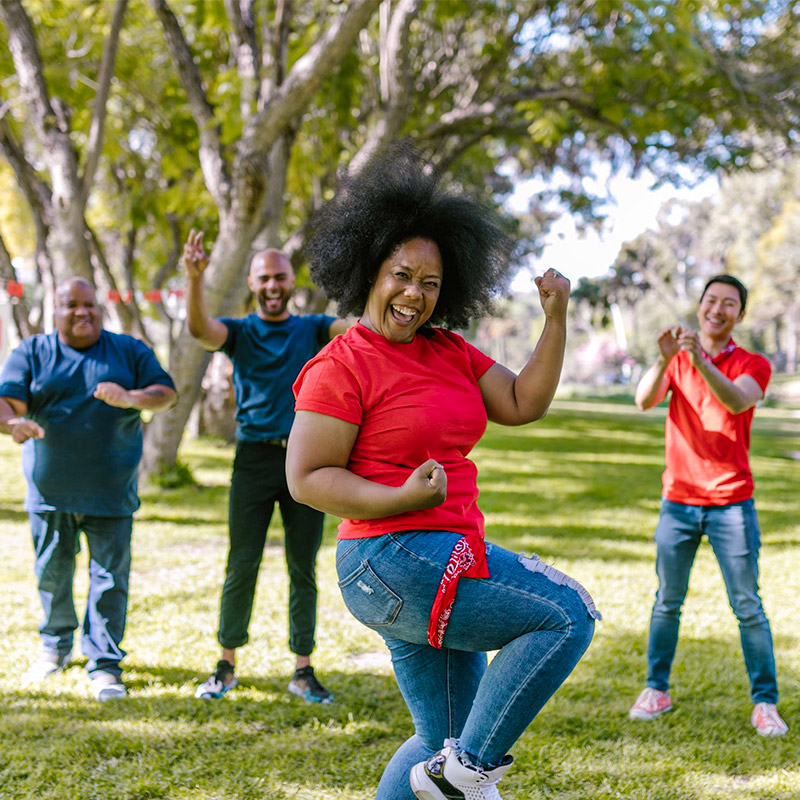

How it works
Step 1
Purchase a Medication kit.
Step 2
Register your kit and nominate your healthcare professional.
Step 3
Swab and send back to the lab using the enclosed envelope.
Step 4
An email will notify you when your results are ready.
Step 5
Book an appointment with your healthcare professional to discuss your results.
If you are eligible for a rebate from your Private Health Insurer, please get a referral from your doctor. You will need a referral, along with a copy of your PGx test receipt when making your claim.
Please note: results will not be released until the consultation with your healthcare practitioner has been undertaken.
Medication Categories
Included in Multiple Category Medication Test Kit.

Mental Health
Find out how your body processes certain medications for depression, ADHD, anxiety and more.

Pain
Find out how your body processes certain anti-inflammatories, opioids and other medications.
Cardiovascular
Find out how your body processes certain cholesterol-lowering medications, beta blockers and other heart medications.

Gastrointestinal
Find out how your body processes certain medications for nausea, reflux, ulcers and more.
The main categories that the Multiple Category Medication Test Covers are highlighted above, however, there are further insights into other medications also included with this test.
To see if we report on the medication that you’ve been prescribed, please contact our friendly Customer Service Team on 1300 436 373.
Benefits of a MYDNA Personalised Medication Test
- Reduce the risk of adverse drug reactions
- Reduce wasted time on ‘trial and error’ medicines
- Reach treatment goals sooner
- Results are relevant for a lifetime
DNA Testing for Medication Tolerance
The study of pharmacogenomics looks at how genes can affect an individual’s response to certain medications. By undertaking the myDNA pharmacogenomic DNA testing for medication, you can be more informed on how your body responds to a range of different medications, such as those used to treat mental health, pain, gastroenterology and cardiology.
Giving You an Insight into How Your Body Responds to Medication
Pharmacogenomic DNA testing for medication response helps your healthcare professional understand the role your genetics plays in how you process medications. This allows them to be better placed to prescribe safer and more effective medications for you.
This knowledge helps to reduce the trial and error approach of prescribing medications so you can receive the right medication faster. Our DNA-based medication insights are available for a range of commonly prescribed medications. Please contact our friendly customer service team to see if we report on the medication that you’ve been prescribed.
Backed by Scientific Evidence and Reviewed by Clinical Experts
Our clinical team has over 120 years of combined clinical experience which they use together with the latest published guidelines and data to create your DNA insights. This helps you and your healthcare professional make more informed choices regarding your medication and dose.
It’s easy to undergo pharmacogenomic DNA medication testing in Australia with myDNA. Simply choose the appropriate testing kit, carry out your cheek swab, and send it to the myDNA Melbourne based lab. Let us do the rest.
Your DNA and your data will always be safe with us. We use secure encrypted servers; we’ll never share your data without your consent and your DNA can be destroyed at any time at your request.
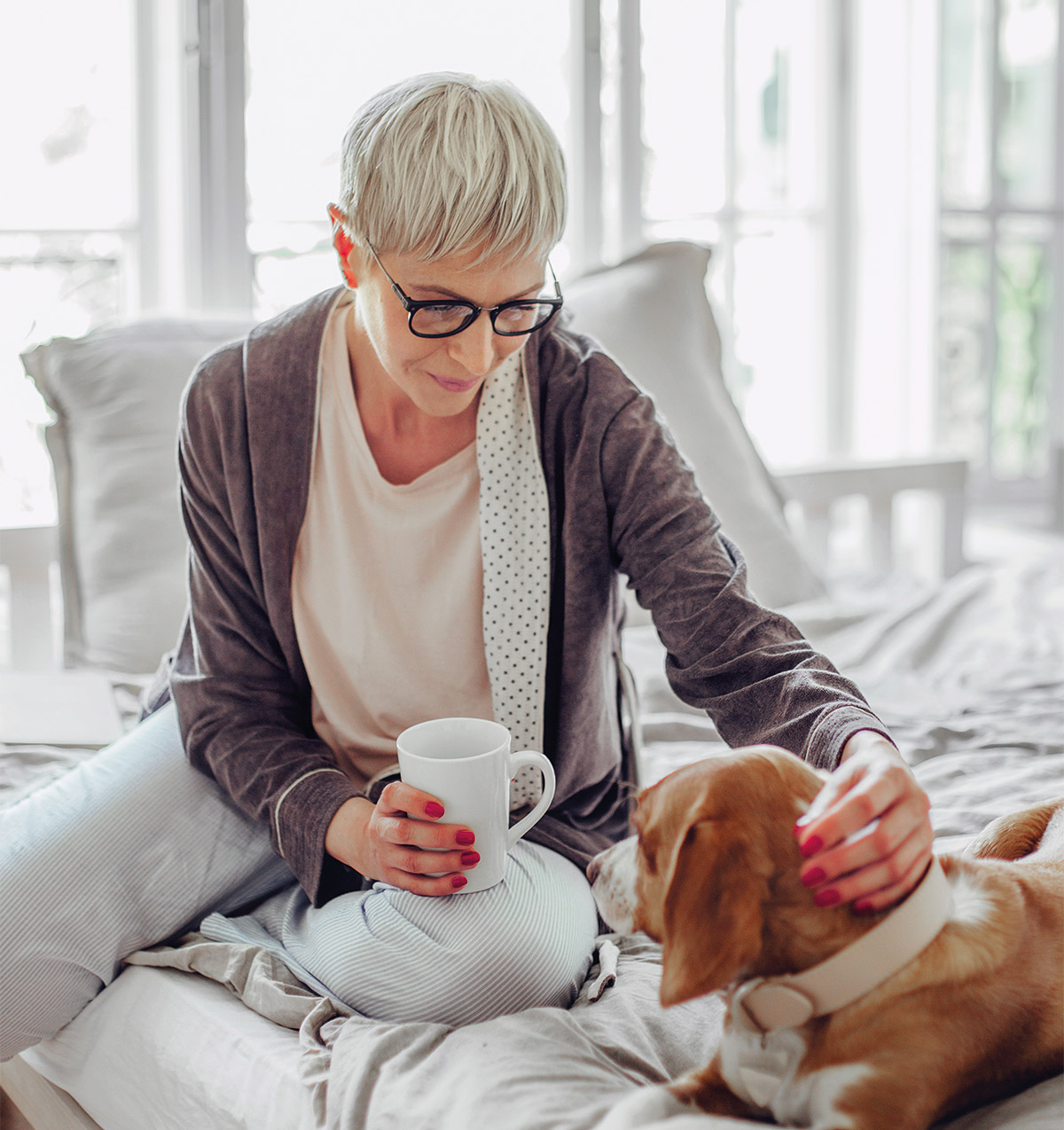
Customer success stories
Rachel Griffith
It’s given us peace of mind… we just know we’re not messing about with medications.
Joy Mawbey
It was more or less a revelation… that there is a reason that I’m suffering from so much pain post surgery compared to others.
Did you Know?
Over one million people in Australia are prescribed antidepressants.
Finding an antidepressant that works is some times a matter of trial-and-error. Only 50% of people respond to the first antidepressant they try.
A simple myDNA Medications Test can help your doctor to tailor your medications to suit your genetic profile.


Your data, your property.

Privacy and data security protocols are fundamental to the myDNA technology platform.
myDNA will interpret and provide a personalised report for the genetic myDNA test requested by you or your healthcare professional only. Any results generated remain strictly confidential and will not be shared with any third parties without your consent.
For more, please read our privacy policy.




